Introducing Textual
I've started work on a new project called Textual.
Textual is (or will be) a TUI framework using Rich as a renderer. It's still very early days, but ultimately the goal is to have something which you can uses to build a Text User Interface with little to no boiler-plate.
It is async powered. Each "widget" runs in an async task and communicates via a message queue. This should allow the UI to remain responsive even when the app is doing work.
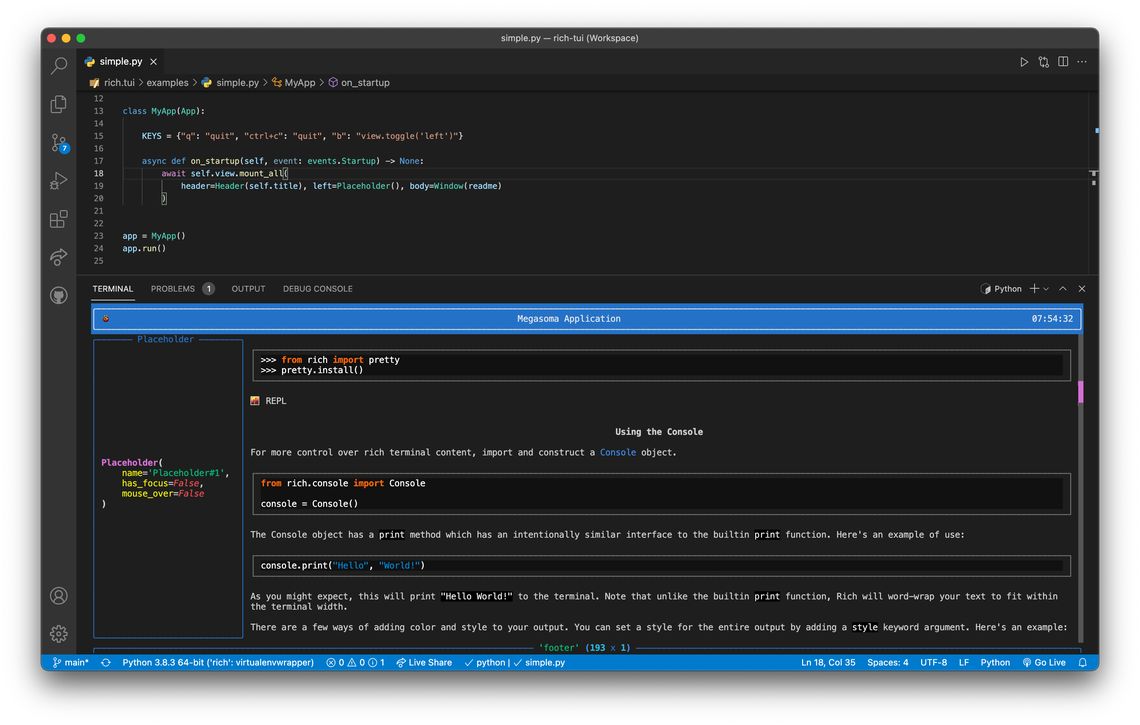
Running a Textual app in Vscode.
An example Textual app (currently the only one) with a default layout. Title bar with working clock. Side panel that can be toggled with a key. And a scrolling window containing Markdown.
Feel free to check out the code, but since Textual is very much a work in progress, you can expect glitches. I'm going to document progress here.
Update 1
Mouse hover, focus, and scrolling.
Update 2
Added key bindings and actions. Video demontrates binding a key to toggle a widget in the layout.
Update 3
Fixed slow scrolling. It turns out that I was parsing and rendering the Markdown every refresh. Not that the cache works, scrolling is much smoother.
Also implemented a new scrollbar using block characters for more granularity.
I was looking for something like this! Super excited!! I was quite close to learn Rust because of tui-rs. :D
'rich' url leads to textual
fixed, thanks.
This looks incredible!
I've been playing with Textual a bit and I'm very excited for the future.
I've been playing with Textual and am excited for more development.